Research Areas
We are developing novel single-cell and spatial multi-omic technologies, together with advanced computational methods to push the boundaries of what can be measured and understood in tissues. We use these methods to study diverse areas of biology such as adipose tissue dysfunction in diabetes, immuno-oncology, and cancer metastasis & therapy resistance.
Next-generation single-cell and spatial multi-omics
Single-cell omics encompasses a rapidly evolving suite of technologies that enable comprehensive molecular characterization of individual cells at high throughput. Spatially resolved omic techniques extend this capability by retaining the native tissue architecture, while multi-omic approaches integrate measurement of several omic features within the same cell. Our lab is focused on advancing the next generation of such technologies, by developing systems that unify single-cell resolution, spatial context, and multi-omic measurements within a single experimental framework. To achieve this, we are developing modular and adaptable assays that can interface with and extend existing commercial platforms, enabling them to access new modes of measurement. In parallel, we also systematically probe the design and performance limits of commercial assays to engineer cost-efficient adaptations. Through these efforts, our goal is to democratize access to advanced, high-content cellular measurements while pushing the conceptual boundaries of what single-cell and spatial omics can reveal.
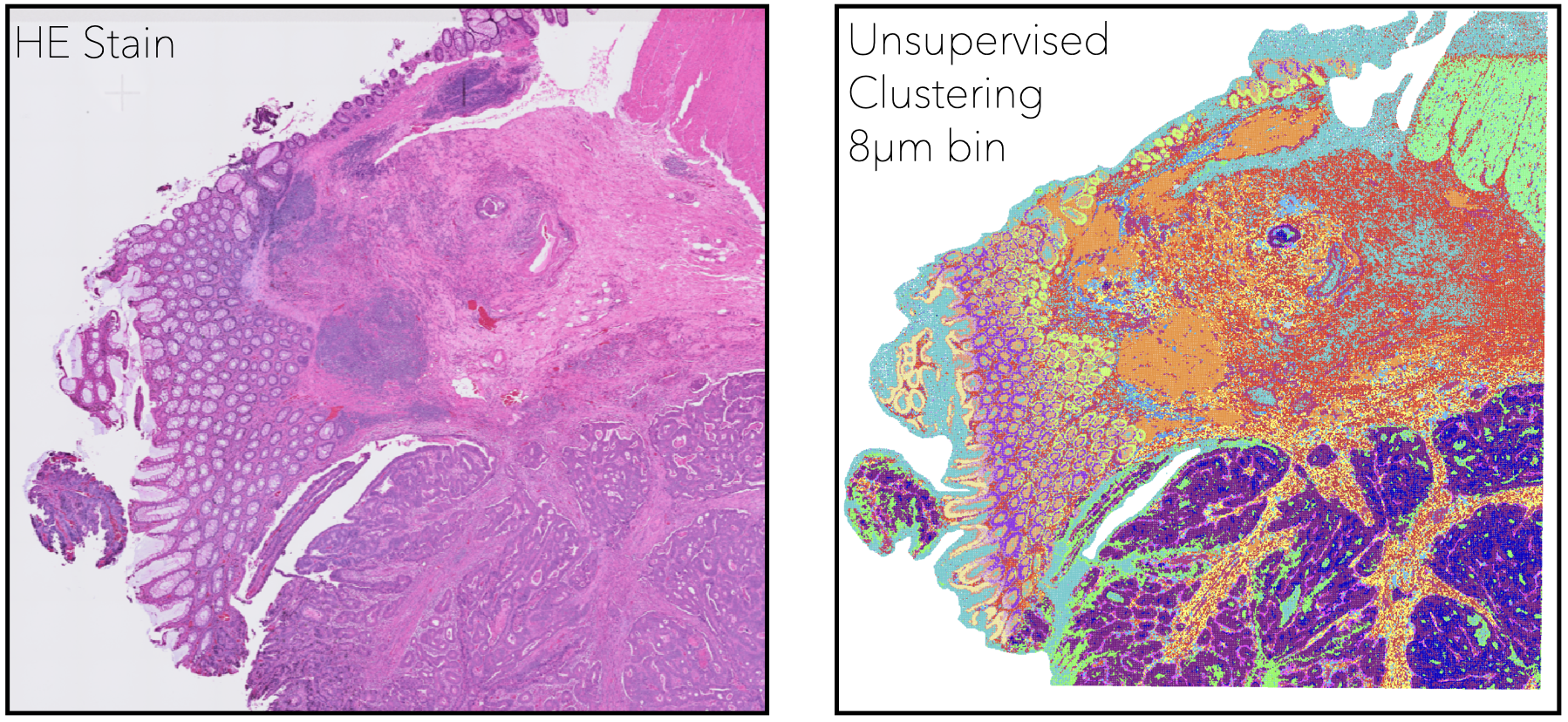
To learn more about our work in this area, see our publications on a microfluidic device for linking transcriptomic and phenomic measurements in single cells, and the development of a novel spatial transcriptomic platform with single-cell resolution.
Bioinformatic methods for single-cell and spatial multi-omics
To process and interpret the massive, high-dimensional datasets generated by single-cell and spatial multi-omic technologies, a rapidly evolving ecosystem of computational methods has emerged, each tailored to specific analytical tasks. However, the performance and applicability of these approaches depend strongly on the biological question, tissue context, omic modality, assay chemistry, and study design, and no single end-to-end pipeline outperforms across all contexts. In our lab, we develop flexible, study-specific bioinformatic workflows that explicitly account for the biological, statistical, and technical characteristics of each dataset. We achieve this by systematically integrating, adapting, and extending existing methods into modular, well-validated pipelines, and by developing new computational approaches when current tools are insufficient to capture system-specific features. Our goal is to deliver robust, interpretable, and reproducible analyses that uncover cell states, spatial organization, regulatory relationships, and coordinated multi-omic programs within complex biological systems.
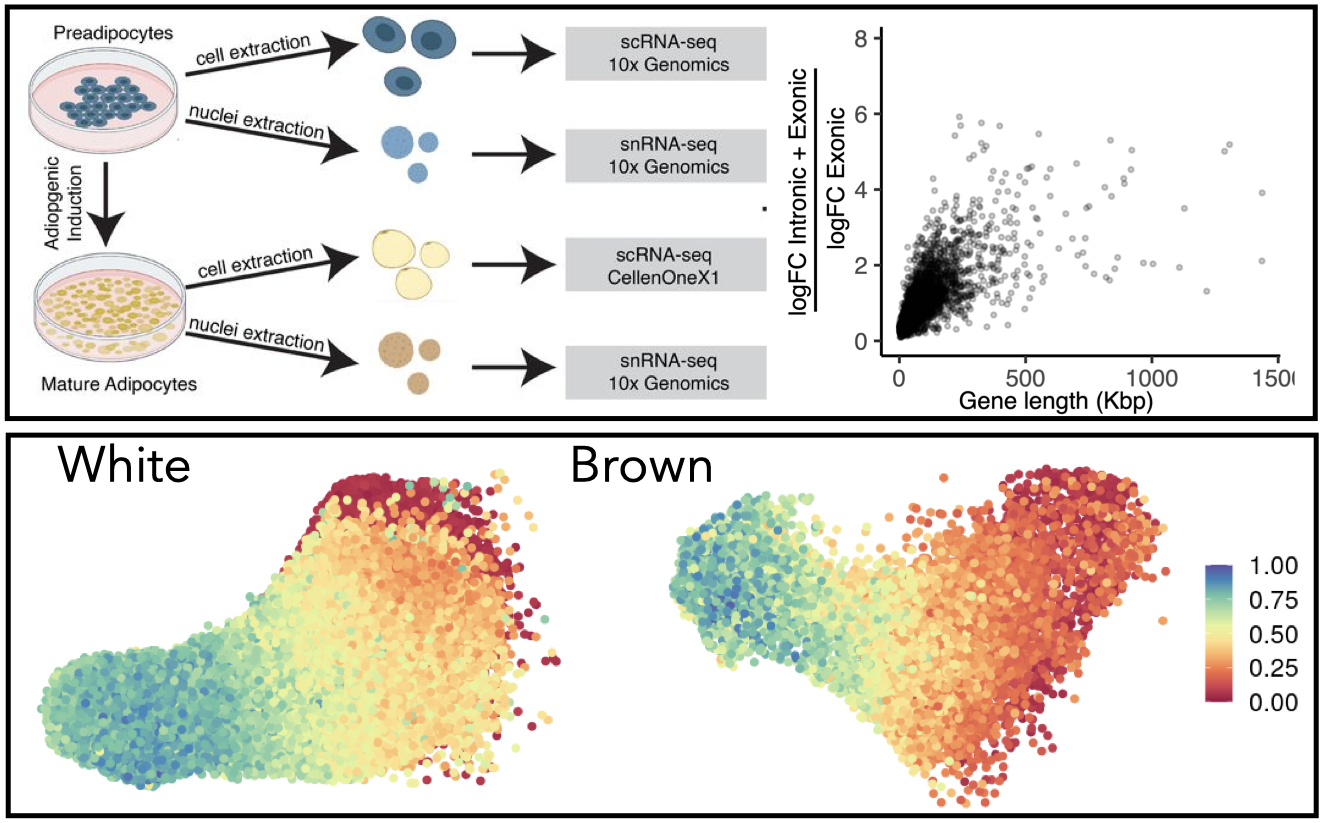
To learn more about our work in this area, see our studies on transcript enrichment and detection bias in single-nucleus versus whole-cell RNA-seq, and on custom pseudotemporal ordering strategies for adipose tissue biology.
Tumor-microenvironmental heterogeneity and their co-evolution during cancer progression
Cancer is a heterogeneous disease where multiple tumor cell states coexist within the same tumor. These states often arise through stepwise genetic diversification, and are further shaped by dynamic interactions with the tumor microenvironment, whose composition and function are in turn remodeled by the tumor cells themselves. Together, what emerges is a spatially and temporally evolving ecosystem, ultimately selecting for cellular states that can invade, metastasize, or resist therapy. In our lab, using our developed single-cell and spatial multi-omic technologies, we are interested in comprehensively mapping the molecular states of tumor and microenvironmental cells, their spatial organization, and understand how these features evolve over time across clinical trajectories including normal tissue to precancerous lesions to invasive disease progression, pre- to post-treatment remodeling, and between primary tumors and secondary metastatic locations. Through these measurements, we aim to build mechanistic models of how heterogeneous tumor ecosystems emerge, adapt, and ultimately drive disease progression and treatment failure.
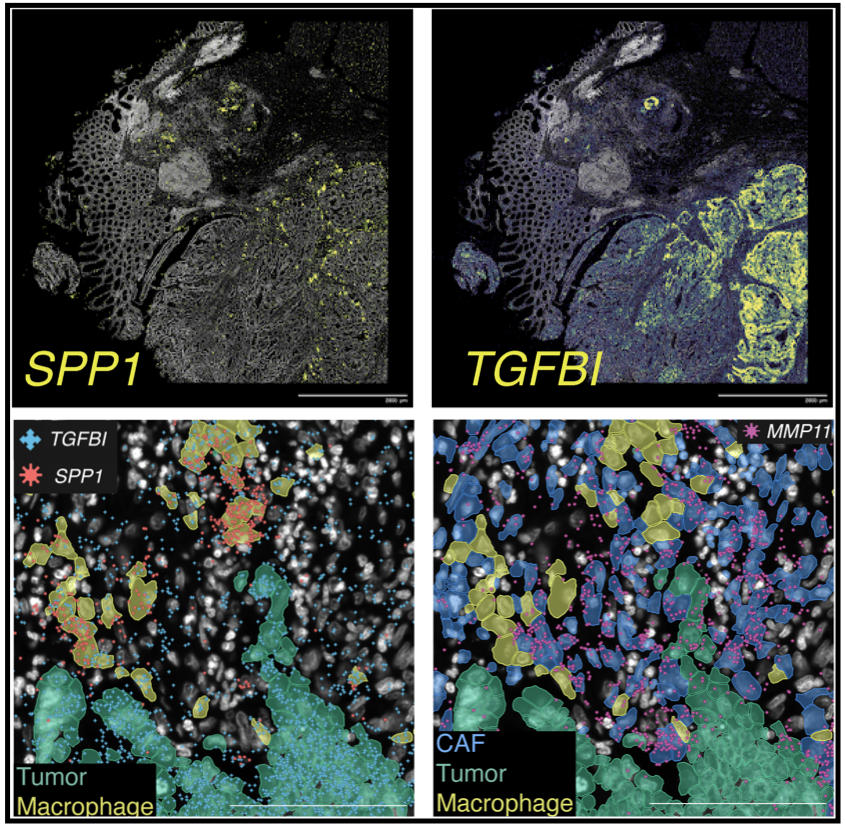
To learn more about our work, read our paper on spatial transcriptomic profiling in colorectal cancer to comprehensively map the landscape of immune cell infiltration and tumour interactions.
Broader applications in biomedical research
Advances in single-cell and spatial omic technologies have reshaped how we investigate biological complexity across development, physiology, and disease. Using our developed multi-omic tools, we aim to push the conceptual boundaries of what single-cell and spatial omics can reveal and contribute to diverse areas of biology. If you have a biological question where spatial context, multimodal measurements, and integrative analysis are essential for new insights, please reach out to us at agupta@iitm.ac.in!
